EcoCyc E. coli Database
EcoCyc captures information from 45,000 publications for Escherichia coli K-12 substr. MG1655. Our two minute tutorials will teach you how to use EcoCyc to search for knowledge on E. coli genes, regulation, and metabolism; analyze transcriptomics data; and perform comparative analyses.New to EcoCyc? See the EcoCyc project overview.
EcoCyc is part of the larger BioCyc collection of thousands of Pathway/Genome Databases for sequenced genomes. Click "Change Current Database" above to explore the available databases.
What people are saying...
Learning Library
Tutorial Videos
Tutorial #1: Introduction to BioCyc
Tutorial #2: Introduction to SmartTables
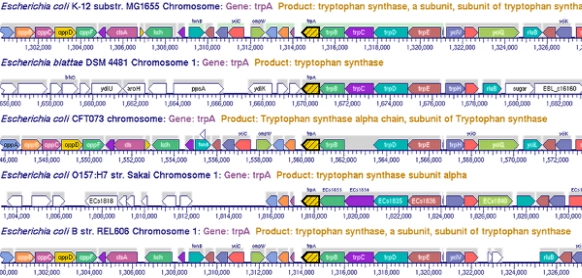
Tutorial #3: Zoomable Metabolic Map, Comparative Tools, Regulatory Network
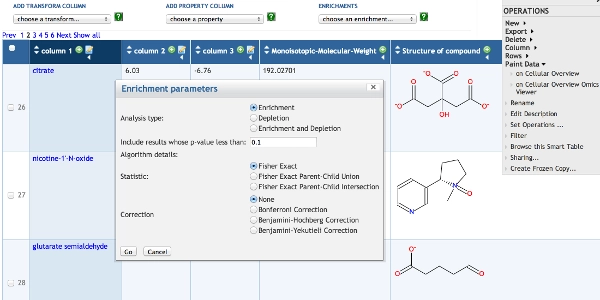
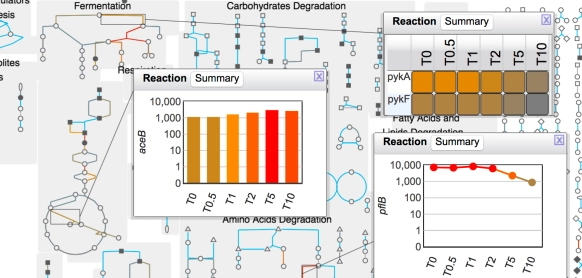
Tutorial #4: Omics Data Analysis
Tutorial #5: Pathway Collages
Tutorial #6: Creating a Pathway/Genome Database
- Part 1A: Introduction to Database Building and Pathologic (14:04)
- Part 1B: Building a Database: Detailed Pathologic Example (23:53)
- Part 2A: General Editing Strategies (8:00)
- Part 2B: Creating and Editing Reactions and Compounds (17:32)
- Part 2C: Updating Proteins, Citations, GO Terms, and Enzymatic Reactions (26:10)
- Part 2D: Making and Editing Pathways (9:42)