EcoCyc E. coli Database
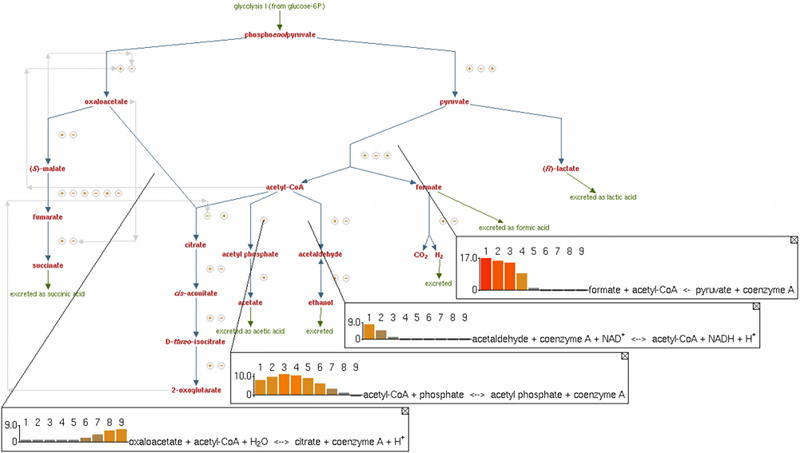
EcoCyc captures information from 44,000 publications for Escherichia coli K-12 substr. MG1655. Use EcoCyc to search for knowledge on E. coli genes, regulation, and metabolism; analyze transcriptomics data; perform comparative analyses; and run a metabolic model.New to EcoCyc? See the EcoCyc project overview.
EcoCyc is part of the larger BioCyc collection of thousands of Pathway/Genome Databases for sequenced genomes. Click "Change Current Database" above to explore the available databases.
What people are saying
"BsubCyc is a tool of the utmost value."

Paul Babitzke
Prof. of Biochemistry
& Molecular Biology
"My lab uses these resources on a daily basis."

Patricia Kiley,
Professor and Chair,
Dep't. of Biomolecular Chemistry
"We rely on BioCyc's Gene Pages and Overview Diagrams almost daily."

Arkady Khodursky
Assoc. Prof. Biochemistry
"We use BioCyc and MetaCyc extensively to investigate the metabolic and regulatory processes of organisms we study."

William Cannon, Team Lead
Computational Biology
"BioCyc is the go-to resource of knowledge and tools for Ginkgo scientists."

"BioCyc is a tremendous resource for pathway analysis in metabolomics."

Art Edison, Dept of Genetics
"We make extensive use of the BioCyc full metabolic network diagram for omics data analysis."

Timothy J. Donohue, Director
"I have not found another database that has a better interface than BioCyc."

Gary B. Huffnagle, Professor
Microbiology and Immunology
Learning Library
Tutorial Series
Tutorial #1: Introduction to BioCyc
Tutorial #2: Introduction to SmartTables
Tutorial #3: Zoomable Metabolic Map, Comparative Tools, Regulatory Network
Tutorial #4: Omics Data Analysis
Tutorial #5: Pathway Collages
Tutorial #6: Creating a Pathway/Genome Database
- Part 1A: Introduction to Database Building and Pathologic (14:04)
- Part 1B: Building a Database: Detailed Pathologic Example (23:53)
- Part 2A: General Editing Strategies (8:00)
- Part 2B: Creating and Editing Reactions and Compounds (17:32)
- Part 2C: Updating Proteins, Citations, GO Terms, and Enzymatic Reactions (26:10)
- Part 2D: Making and Editing Pathways (9:42)